sparkConf": {
"spark.eventLog.enabled": "true",
"spark.network.timeout": "300s",
"spark.task.maxFailures": "10",
"spark.executor.memory": "22g",
"spark.executor.cores": "2",
"spark.driver.memory": "100g",
"spark.driver.maxResultSize": "20g",
"spark.driver.cores" : "16",
"spark.hadoop.fs.s3n.multipart.uploads.split.size": "536870912",
"spark.yarn.executor.memoryOverheadFactor": "0.20",
"spark.yarn.driver.memoryOverheadFactor": "0.20",
"spark.shuffle.registration.timeout": "30s",
"spark.shuffle.sasl.timeout": "120s",
"spark.sql.shuffle.partitions": "30000",
"spark.hadoop.mapreduce.fileoutputcommitter.algorithm.version": "2",
"spark.network.auth.rpcTimeout":"180s",
"spark.executor.heartbeatInterval":"30s",
"spark.yarn.am.memory":"100g",
"spark.yarn.am.cores":"16",
"spark.reducer.maxReqsInFlight":"1",
"spark.shuffle.io.retryWait":"20s",
"spark.shuffle.io.maxRetries":"5",
"spark.shuffle.compress":"true",
"spark.shuffle.spill.compress":"true",
"spark.unsafe.sorter.spill.read.ahead.enabled" : "false",
"spark.rdd.compress":"true",
"spark.checkpoint.compress":"true",
"spark.broadcast.compress":"true",
"spark.sql.inMemoryColumnarStorage.compressed":"true",
"spark.speculation":"true",
"spark.speculation.quantile":"0.85",
"spark.speculation.multiplier":"3",
"spark.scheduler.listenerbus.eventqueue.capacity":"20000"
}Author: eulertech
How to control spark disk output file size and number of part files
While processing big volume of data in spark, the data in the memory is partitioned per spark.sql.shuffle.partitions numbers which can be set by (sqlContext.setConf("spark.sql.shuffle.partitions", "300")).This number of partition will be used when shuffling for joining or aggregation. This number will also impact the number of part files being generated while spark.write to disk. Another magic number is 2000, which spark have different logic for data compression (>2000 has data compressed). If we serialize the data from memory to the disk using all default, we’re going to a lot files potentially many small files. In this article, I’ll show the scenarios and how to control the part file size and total number of files in each partition or folder.
In this article, the default shuffle partition is equal to 2.
For demo the situation, a random data generator is created to simulate skewed data situation. This will generate data per country with varying size. The data will look like this:
+—+—-+——-+
|id |sub |country|
+—+—-+——-+
|0 |Even|Fiji |
|1 |Odd |Fiji |
|2 |Even|Fiji |
|3 |Even|Iceland|
|4 |Odd |Iceland|
|5 |Even|Iceland|
|6 |Odd |Iceland|
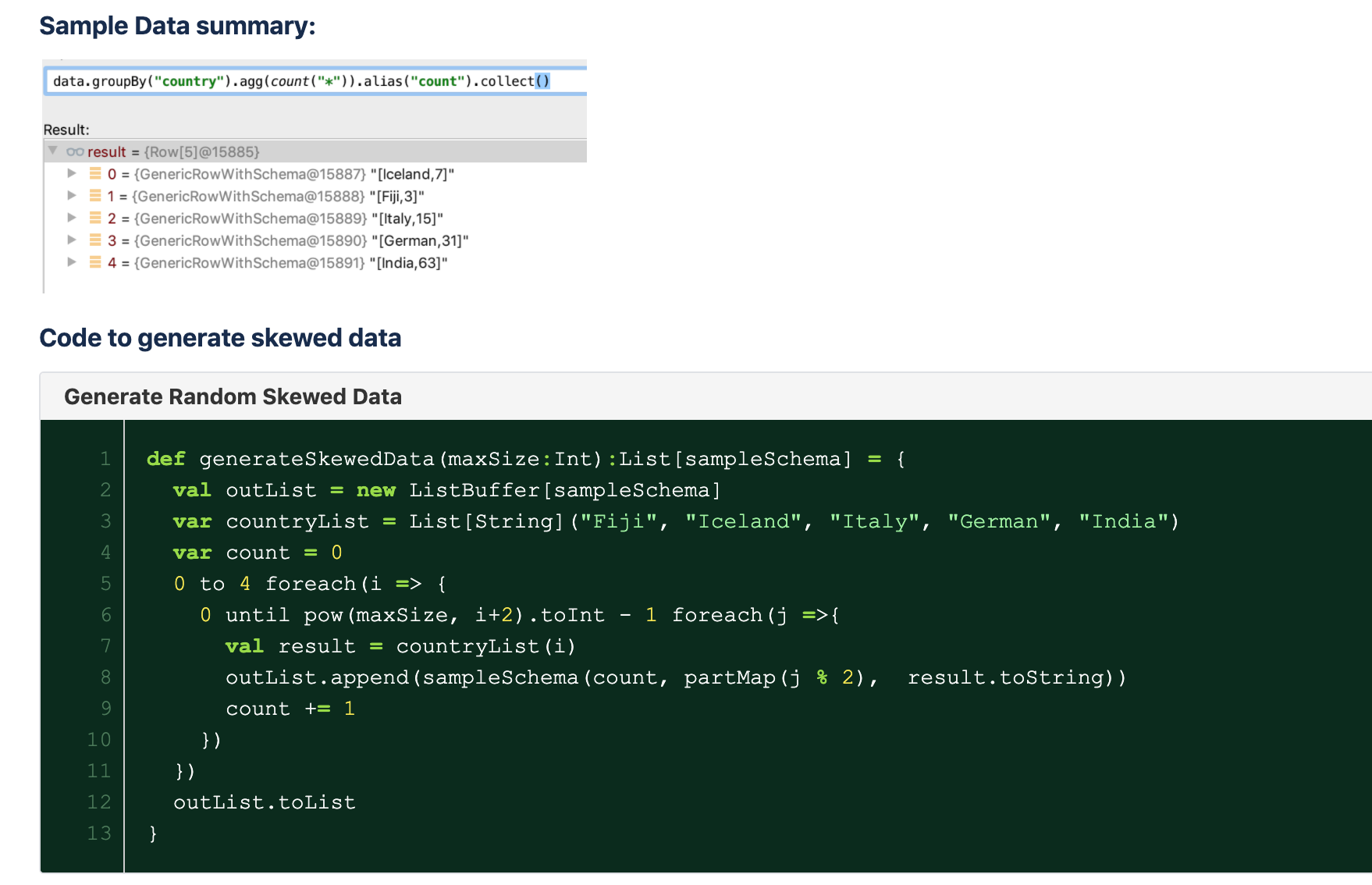








In summary, we can simple use option(“maxRecordsPerFile”) and partitionBy to control the number of records in each part file. To remove small and big file problem. Data after repartition by column will evenly re-distribute and the number of files is dynamically generated per data sample.
Simple RegExp to preprocess text
While doing with NLP projects, the first step is to always clean the data. Mostly commonly remove all non-alphanumeric, HTML tags, emojis, etc. Then we’ll move to stemming, lemmatization, n-grams or word-embedding, etc.
Here is a summary of the pre-cleaning text using pure regex in Python:
</pre>
<pre>def remove_non_alphanum(text):
import re
clean = re.sub(r'\W+', '', text)
return clean
def remove_html_tags(text):
"""Remove html tags from a string"""
import re
clean = re.compile('')
return re.sub(clean, '', text)</pre>
<pre>
Remove emojis:
</pre>
<pre>text = u'This dog \U0001f602'
print(text) # with emoji
def remove_emoji(text):
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
"]+", flags=re.UNICODE)
return emoji_pattern.sub(r'', text) # no emoji</pre>
<pre>
Python libraries for commonly text search and comparison tasks
All of us are familiar with searching a text for a specified word or character sequence (pattern). The goal is to either find the exact occurrence (match) or to find an in-exact match using characters with a special meaning, for example by regular expressions or by fuzzy logic. Mostly, it is a sequence of characters that is similar to another one.
Furthermore, the similarity can be measured by the way words sound — do they sound similar but are written in a different way? Translations from one alphabet to another often gives more than one result depending on the language, so to find relatives based on the different spellings of their surname and name the Soundex algorithm was created and is still one of the most popular and widespread ones today.
Table 1 covers a selection of ways to search and compare text data. The right column of the table contains a selection of the corresponding Python modules to achieve these tasks. (ref: https://stackabuse.com/levenshtein-distance-and-text-similarity-in-python/)
| Category | Method or Algorithm | Python packages |
|---|---|---|
| Exact search | Boyer-Moore string search, Rabin-Karp string search, Knuth-Morris-Pratt (KMP), Regular Expressions | string, re, Advas |
| In-exact search | bigram search, trigram search, fuzzy logic | Fuzzy |
| Phonetic algorithms | Soundex, Metaphone, Double Metaphone, Caverphone, NYIIS, Kölner Phonetik, Match Rating codex | Advas, Fuzzy, jellyfish, phonetics, kph |
| Changes or edits | Levenshtein distance, Hamming distance, Jaro distance, Jaro-Winkler distance | editdistance, python-Levenshtein, jellyfish |
Create an in-memory database with SQLite in Python
An in-memory database can improve the performance while a lot of reading/writing processes occurred. In-memory databases are most commonly used in applications that demand very fast data access, storage and manipulation, and in systems that don’t typically have a disk but nevertheless must manage appreciable quantities of data. An important use for in-memory database systems is in real-time embedded systems.
What’s the best way to create an in-memory database with python?
Step : create the connection outside the db_execute function or create a shared memory connection:
A plain :memory: the string connecting to an in-memory database cannot be shared or attached to from other connections.
Keep in mind that the database will be erased when the last connection is deleted from memory. A.K.A connection closed.
def execute_db(args):
try:
with db:
cur = db.cursor()
cur.execute(*args)
return true
except Exception as why:
return False
The transaction is automatically committed if there was no exception, rolled back otherwise. Note that it is safe to commit a query that only reads data.
non-static method can’t be referenced in static context
While trying to call a Scala singleton method from Java, you might have seen errors like: non-static method [method name] can’t be referenced in static context. What does it mean and how to fix it?
First, as a top-level value, an object is a singleton. And an object is a class that has exactly one instance. Scala has a unique way of create singleton with companion object for each class. What is companion object: An object with the same name as a class is called a companion object. Example:
import scala.math._
class Circle(radius: Double) {
override def name:String = “name” + radius
}
object Circle {
def name:String = “object name”
}
In Scala, you can directly call Circle.name to get “object name”.
However, when you want to do same thing in Java, it will give you above error.
First, how to reference a singleton method in Java? Simple, Circle$.MODULE$.name()
Why?
Scala differentiates namespaces between Object and Class/Trait. They can use the same name. However, an object has a class, and therefore needs an angled name on the JVM. The current Scala conventions is to add a $ at the end of the module name. If the object is defined in a class, its convention is OuterClass$ModuleName$. In 2.8.0 this means if you have a companion object, you lose your static forwarders.
In this case a “best practice” would be to always use the Circle$.MODULE$ reference instead of a static forwarder, as the forwarder could disappear with modifications to the Circle class.
Pycharm console can’t connect
Have you ever encountered issues while trying to start python console within Pycharm but it gives you an error like this:

It is a little annoying when issues like this happen. For quickly dissolve the issue, simply edit the pydev_ipython_console_011.py to make it work. It is due to ipython version 3.6.3.0 bug.
Go to file: “/Applications/PyCharm CE.app/Contents/helpers/pydev/_pydev_bundle/pydev_ipython_console_011.py” Line 85 and make the following change. Restart pycharm and it should work.

Union two Spark dataframes with different schema
Image that you have two dataframes with different schema but there are some common columns too and you want to union these two dataframe together. Let’s look at this example:
import org.apache.spark.sql.functions._
val df1 = sc.parallelize(List((21,”john”),(24,”alex”))).toDF(“age”,”name”)
val df2 = sc.parallelize(List((33,”France”,”Math”),(21,”Italy”,”Physics”))).toDF(“age”,”country”,”major”)
Now the two dataframe has different columns, one has columns of (age, name) while the other has columns of (age, country, major). How do we union them together?
val cols1 = df1.columns.toSet
val cols2 = df2.columns.toSet
val allCols = cols1 ++ cols2
def expr(myCols:: Set[String], allCols: Set[String]) = {
allCols.toList.map(x => x match{
case x if myCols.contains(x) => col(x)
case _ => list(null).as(x)
}
}
df1.select(expr(cols1, total):_*).unionAll(df2.select(expr(cols2, total):_*)).show()
All about date, time parsing with Scala
Time has been the ubiquitous dimension in very dataset we collect, from stock market to weather stations and Facebook conversations history. While the data can be really messy, the data formats stored can be in epoch, various mixed time formats like: 2019-11-17 11:12:00, 2019-11-17 11:12:00.123, 2019-11-17 11:12:00.000000123 or 20191117T111200.000 etc. Also, we need to take timezone into consideration.
Here is the function that I have compiled to parse mixed type of format in one pass:
import java.math.BigDecimal
import java.sql.{Date, Timestamp}
import java.lang.Long
import java.text.SimpleDateFormat
import java.time.{Instant, LocalDate, LocalDateTime, ZoneId}
import java.time.format.{DateTimeFormatter, DateTimeFormatterBuilder}
import java.time.temporal.ChronoField
def parseEpochTime(epoch: Long): Timestamp = {
val DEFAULT_TIMEZONE = "America/New_York"
try{
Timestamp.valueOf(Instant.ofEpochMilli(epoch).atZone(ZoneId.of(DEFAULT_TIMEZONE)).toLocalDateTime)
}catch{
case e:Exception => {
throw new RuntimeException(s"Can't parse epoch time with ${epoch}")
}
}
}
/**
* This method enabled parse mixture timestamp strings
* Covers: epoch time in millisecond (system default), timestamp and timestamp T format
* @param ts
* @return
*/
def parseMixedTimestamp(ts: String): Timestamp = {
val parser1 = new DateTimeFormatterBuilder()
.appendPattern("yyyy-MM-dd HH:mm:ss")
.appendFraction(ChronoField.NANO_OF_SECOND,0,9,true)
.toFormatter()
val parser2 = new DateTimeFormatterBuilder()
.appendPattern("yyyyMMdd'T'HHmmss")
.appendFraction(ChronoField.NANO_OF_SECOND,0,9,true)
.toFormatter()
try {
val parser = new DateTimeFormatterBuilder().appendOptional(parser1).appendOptional(parser2).toFormatter()
val ldt = LocalDateTime.parse(ts, parser)
val zdt = ldt.atZone(ZoneId.of(DEFAULT_TIMEZONE))
Timestamp.from(zdt.toInstant)
} catch {
case e: Exception => {
e.printStackTrace()
parseEpochTime(ts.toLong)}
}
}
/**
* This method will parse a mixture of date format
* @param dateString
* @param listOfDatePattern A list of acceptable date patterns
* @return
*/
def parseMixedDate(dateString: String, listOfDatePattern: List[String] = List(DATE_FORMAT_PATTERN)): Date = {
val dateTimeFormatter:DateTimeFormatter = listOfDatePattern
.map(DateTimeFormatter.ofPattern)
.foldLeft(new DateTimeFormatterBuilder)((a1:DateTimeFormatterBuilder, a2: DateTimeFormatter) => {
a1.appendOptional(a2)
}).toFormatter
val ldt = LocalDate.parse(dateString, dateTimeFormatter)
Date.valueOf(ldt)
}
// Let's look at the unit test.
@Test
def testTimestamp():Unit = {
val ldt = LocalDateTime.of(2019,10,4,15,31,44,0)
val ts1 = Timestamp.from(ldt.atZone(ZoneId.of(DEFAULT_TIMEZONE)).toInstant)
val tsString = "2019-10-04 15:31:44.000"
val parsedTimestamp = parseMixedTimestamp(tsString)
val tsString2 = "20191004T153144.000"
val parsedTimestamp2 = parseMixedTimestamp(tsString2)
val tsString3 = "20191004T153144"
val parsedTimestamp3 = parseMixedTimestamp(tsString3)
assertEquals(ts1, parsedTimestamp)
assertEquals(ts1, parsedTimestamp2)
assertEquals(ts1, parsedTimestamp3)
}
@Test
def testTimestamp5():Unit = {
val ldt = LocalDateTime.of(2019,10,4,15,31,44,100000000)
val ts1 = Timestamp.from(ldt.atZone(ZoneId.of(DEFAULT_TIMEZONE)).toInstant)
val tsString1 = "2019-10-04 15:31:44.1"
val tsString2 = "2019-10-04 15:31:44.10"
val tsString3 = "2019-10-04 15:31:44.100"
val tsString4 = "2019-10-04 15:31:44.1000"
val tsString5 = "2019-10-04 15:31:44.10000"
val tsString6 = "2019-10-04 15:31:44.100000"
val tsString7 = "2019-10-04 15:31:44.1000000"
val tsString8 = "2019-10-04 15:31:44.10000000"
val tsString9 = "2019-10-04 15:31:44.100000000"
val parsedTimestamp1 = parseMixedTimestamp(tsString1)
val parsedTimestamp2 = parseMixedTimestamp(tsString2)
val parsedTimestamp3 = parseMixedTimestamp(tsString3)
val parsedTimestamp4 = parseMixedTimestamp(tsString4)
val parsedTimestamp5 = parseMixedTimestamp(tsString5)
val parsedTimestamp6 = parseMixedTimestamp(tsString6)
val parsedTimestamp7 = parseMixedTimestamp(tsString7)
val parsedTimestamp8 = parseMixedTimestamp(tsString8)
val parsedTimestamp9 = parseMixedTimestamp(tsString9)
assertEquals(ts1, parsedTimestamp1)
assertEquals(ts1, parsedTimestamp2)
assertEquals(ts1, parsedTimestamp3)
assertEquals(ts1, parsedTimestamp4)
assertEquals(ts1, parsedTimestamp5)
assertEquals(ts1, parsedTimestamp6)
assertEquals(ts1, parsedTimestamp7)
assertEquals(ts1, parsedTimestamp8)
assertEquals(ts1, parsedTimestamp9)
}
@Test
def testMixedDate():Unit = {
val ldt = LocalDate.of(2019,10,4)
val dt1 = Date.valueOf(ldt)
val listOfDatePatterns = List("yyyy-MM-dd", "MM/dd/yyyy")
val parsedDate = parseMixedDate("2019-10-04", listOfDatePatterns)
val parsedDate2 = parseMixedDate("10/04/2019", listOfDatePatterns)
assertEquals(dt1, parsedDate)
assertEquals(dt1, parsedDate2)
}
With these utilities, now you can easily parse all kind of format of date, time at various precision all at once.
How to create sub-partition on an external hard drive on Mac Sierra + with CLI
This article is a little off-topic from ML but it is useful to save your time from focusing on interesting stuff. I have bought some external hard drive routinely back my stuff put on Cloud and PC/Mac since I don’t feel comfortable to let the big companies decide when to mess up my files.
With recent Mac OS changes, partition an external hard drive to a mixed Mac and Windows FS with its util disk app is not that obvious. It has Partition greyed out if the hard drive has master boot drive. Here is a simple command to fix the issue: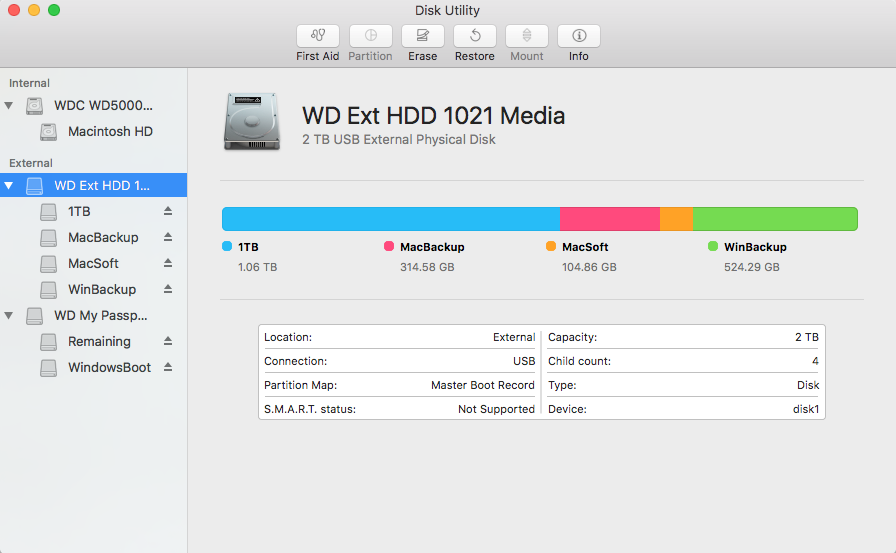
sudo diskutil partitionDisk disk2 2 GPT UFSD_NTFS Remaining R UFSD_NTFS SubPartitoin1 10G
where disk2 is the drive name on your Mac, Purple color is the OS Format (NTFS here) and Blue is the partition names. This example makes two partition, where one is 10 GB, the rest goes to Remaining partition.
Be cautious, this will delete all your data from existing parition!!!!
Before running this command, run diskutil list to list all the drive names.
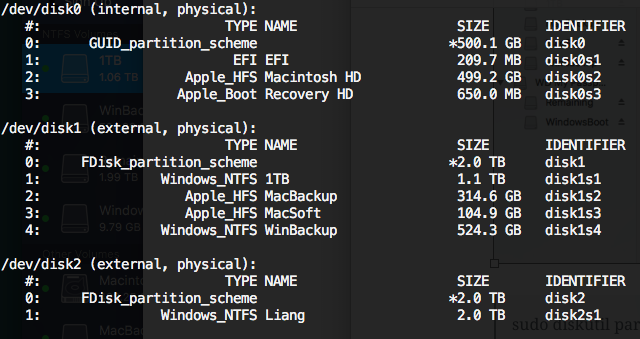
To get all supported file system format name, use: diskutil listFilesystems with the following options: FAT32, HFS+, UFSD_NTFS etc.

